How to detect diseases at the earliest stages of development? This is the problematic raised by most scientists and physicians, focusing on new biomarkers, such a microRNAs. Indeed, a growing number of publications link microRNA levels inside biological liquids to several diseases including some cancers. However, these microRNA molecules are very short, and diluted. Scientists need extremely sensitive techniques to detect and quantify them..
A team from the Gulliver Laboratory at ESPCI Paris (PSL University) worked together with a clinical team in cancerology from the University of Paris, and proposed a solution: a reprogrammable molecular circuit allowing for specifically quantifying microRNA. The discovery has been published lately in Science Advances.

As a world specialist of molecular circuits, Yannick Rondelez has been exploring the biochemichal network reactions to complete complex tasks. He and his team at the Gulliver lab of ESPCI Paris turned a fundamental concept into a molecular circuit capable of detecting, amplifying, and signaling by fluorescence the presence of microRNAs. This test applied to liquid biopsies is based upon a dynamic threshold, which avoids the false positive results.
There is a remarkable analogy with electronics: a first module detects the microRNA for which the circuit was made. The presence is converted into a signal molecule, here a strand of DNA. A second module amplifies exponentially the signal molecule. The third one absorbs the molecular noise, thus reducing the leak effects responsible for false positive results. In the end, the last module produces a light signal after a time that depends on the amount of signal molecules. Recording the fluorescence is a way to go up to the initial concentration of microRNA inside the sample.
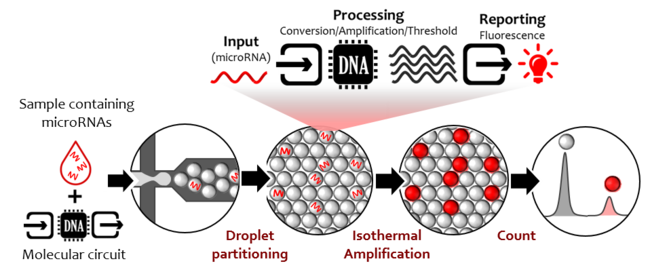
The team also developed a second “digital” approach, to isolate and detect single microRNA molecules. It consists in partitioning the mixture sample/molecular circuit in tiny droplets, such that each one of them will statistically contain only 0 or 1 microRNA molecule. The reaction inside each droplet will « turn on » the filled ones, and the counting of fluorescent droplets is a way to absolutely quantify the microRNA concentration.
These tests have been realized from microRNA samples from human colons, so in close to real conditions. In these cases, microRNA concentrations are minute quantities, between 10-12 (picomolar) and 10-15 (femtomolar) mol/L.
« This technical proof of concept is very promising and the next step will be to validate our method in a clinical frame, with liquid biopsies from patients » explains Guillaume Gines, postdoctoral researcher at ESPCI Paris and co-author of the publication.
The technology is also easily re-programmable to adapt tothe detection to any interesting microRNA, and has also been patented.
« We paved the way for measuring different microRNAs quantities at once, thus hoping for stronger and more precise diagnosis. We are also planning to use the technique on other biomarkers with strong therapeutic interest, like enzymes or proteins » says Yannick Rondelez.
Some actual applications then, that come to conclude years of fundamental research.
Reference :
Gines, Menezes et al.,
Isothermal digital detection of microRNAs using background-free molecular circuit, Science Advances, 2020 (DOI: 10.1126/sciadv.aay5952)
See also : Menezes et al. Streamlined digital bioassays with a 3D printed sample changer, Analyst, 2020 (https://doi.org/10.1039/C9AN01744E)

